Chapter 6 Enrichment analysis
TCMDATA provides functions for performing enrichment analysis on gene sets derived from herb–target interactions. It supports both Over-Representation Analysis (ORA) and Gene Set Enrichment Analysis (GSEA) methodologies via clusterProfiler, and offers a suite of visualization methods for interpreting enrichment results through enrichplot and other compatible packages.
6.1 Over Representation Analysis (ORA)
ORA evaluates whether a pre-defined gene set (e.g., KEGG pathways, GO terms, Reactome pathways) is statistically over-represented among a list of input genes, such as the intersection of DEGs and disease targets. For a comprehensive treatment of enrichment methods, refer to the Biomedical Knowledge Mining book.
6.1.1 KEGG enrichment analysis
KEGG (Kyoto Encyclopedia of Genes and Genomes) is a comprehensive database that maps genes to metabolic pathways, signaling cascades, and other molecular interaction networks. KEGG pathway enrichment analysis identifies which biological pathways are significantly over-represented in a given gene set, thereby revealing the functional mechanisms underlying the observed gene expression changes.
In this example, we randomly sampled 200 targets of Lingzhi (Ganoderma lucidum) as input for KEGG pathway enrichment. Since enrichKEGG() requires Entrez IDs by default, we first converted gene symbols using bitr():
library(TCMDATA)
set.seed(2026)
lz_targets <- search_herb("lingzhi", type = "Herb_pinyin_name")$target
query_genes <- sample(lz_targets, 200)
head(query_genes)#> [1] "YES1" "BCHE" "CYP3A4" "STAT3" "MMP3" "PRKACA"library(clusterProfiler)
library(org.Hs.eg.db)
## convert Symbol to Entrez ID
gene_id <- bitr(query_genes,
fromType = "SYMBOL",
toType = "ENTREZID",
OrgDb = org.Hs.eg.db)
head(gene_id)#> SYMBOL ENTREZID
#> 1 YES1 7525
#> 2 BCHE 590
#> 3 CYP3A4 1576
#> 4 STAT3 6774
#> 5 MMP3 4314
#> 6 PRKACA 5566Then, we performed KEGG pathway enrichment using enrichKEGG(), and applied setReadable() to convert enriched gene IDs back to symbols for better interpretability:
kegg_enrich <- enrichKEGG(gene = gene_id$ENTREZID,
organism = "hsa",
keyType = "kegg")
kegg_enrich <- setReadable(kegg_enrich, OrgDb = org.Hs.eg.db, keyType = "ENTREZID")
head(kegg_enrich)#> category subcategory ID
#> hsa05160 Human Diseases Infectious disease: viral hsa05160
#> hsa04657 Organismal Systems Immune system hsa04657
#> hsa05219 Human Diseases Cancer: specific types hsa05219
#> hsa05417 Human Diseases Cardiovascular disease hsa05417
#> hsa01522 Human Diseases Drug resistance: antineoplastic hsa01522
#> hsa00910 Metabolism Energy metabolism hsa00910
#> Description GeneRatio BgRatio RichFactor FoldEnrichment
#> hsa05160 Hepatitis C 20/156 156/9260 0.12820513 7.610125
#> hsa04657 IL-17 signaling pathway 15/156 94/9260 0.15957447 9.472177
#> hsa05219 Bladder cancer 11/156 41/9260 0.26829268 15.925578
#> hsa05417 Lipid and atherosclerosis 21/156 216/9260 0.09722222 5.771011
#> hsa01522 Endocrine resistance 15/156 99/9260 0.15151515 8.993784
#> hsa00910 Nitrogen metabolism 8/156 17/9260 0.47058824 27.933635
#> zScore pvalue p.adjust qvalue
#> hsa05160 10.89893 1.182718e-12 3.311610e-10 1.232517e-10
#> hsa04657 10.80680 3.750864e-11 4.155521e-09 1.546604e-09
#> hsa05219 12.53746 4.452344e-11 4.155521e-09 1.546604e-09
#> hsa05417 9.28721 6.848868e-11 4.527151e-09 1.684917e-09
#> hsa01522 10.46711 8.084198e-11 4.527151e-09 1.684917e-09
#> hsa00910 14.54927 1.158830e-10 4.701066e-09 1.749645e-09
#> geneID
#> hsa05160 STAT3/MAPK1/CYCS/PIK3CD/EGFR/GSK3B/PPARA/TNF/TRAF6/CDKN1A/TP53/BRAF/CASP3/AKT2/CDK4/CFLAR/STAT2/NFKBIA/E2F1/FADD
#> hsa04657 MMP3/MAPK1/IL4/GSK3B/TNF/TRAF6/MMP9/CCL2/MMP1/CASP3/FOS/TNFAIP3/IL17A/NFKBIA/FADD
#> hsa05219 MAPK1/ERBB2/EGFR/CDKN1A/TP53/MMP9/BRAF/MMP1/CDK4/DAPK1/E2F1
#> hsa05417 STAT3/MMP3/MAPK1/CYCS/PIK3CD/GSK3B/TNF/CAMK2B/TRAF6/TP53/CASP1/MMP9/CCL2/MMP1/CASP3/AKT2/CYP1A1/FOS/CYP2C8/PYCARD/NFKBIA
#> hsa01522 PRKACA/ADCY1/MAPK1/ERBB2/PIK3CD/EGFR/CDKN1A/TP53/MMP9/BRAF/AKT2/CDK4/FOS/IGF1R/E2F1
#> hsa00910 CA2/CA4/CA5A/CA5B/CA7/CA14/CA13/CA9
#> Count
#> hsa05160 20
#> hsa04657 15
#> hsa05219 11
#> hsa05417 21
#> hsa01522 15
#> hsa00910 8The enrichplot package is fully compatible with clusterProfiler and provides a rich set of visualization methods. Below we demonstrated several common plot types (dot plot, bar plot, and concept network plot):
p1 <- dotplot(kegg_enrich)
p2 <- barplot(kegg_enrich)
p3 <- cnetplot(kegg_enrich)
plot_list(p1, p2, p3, ncol = 1)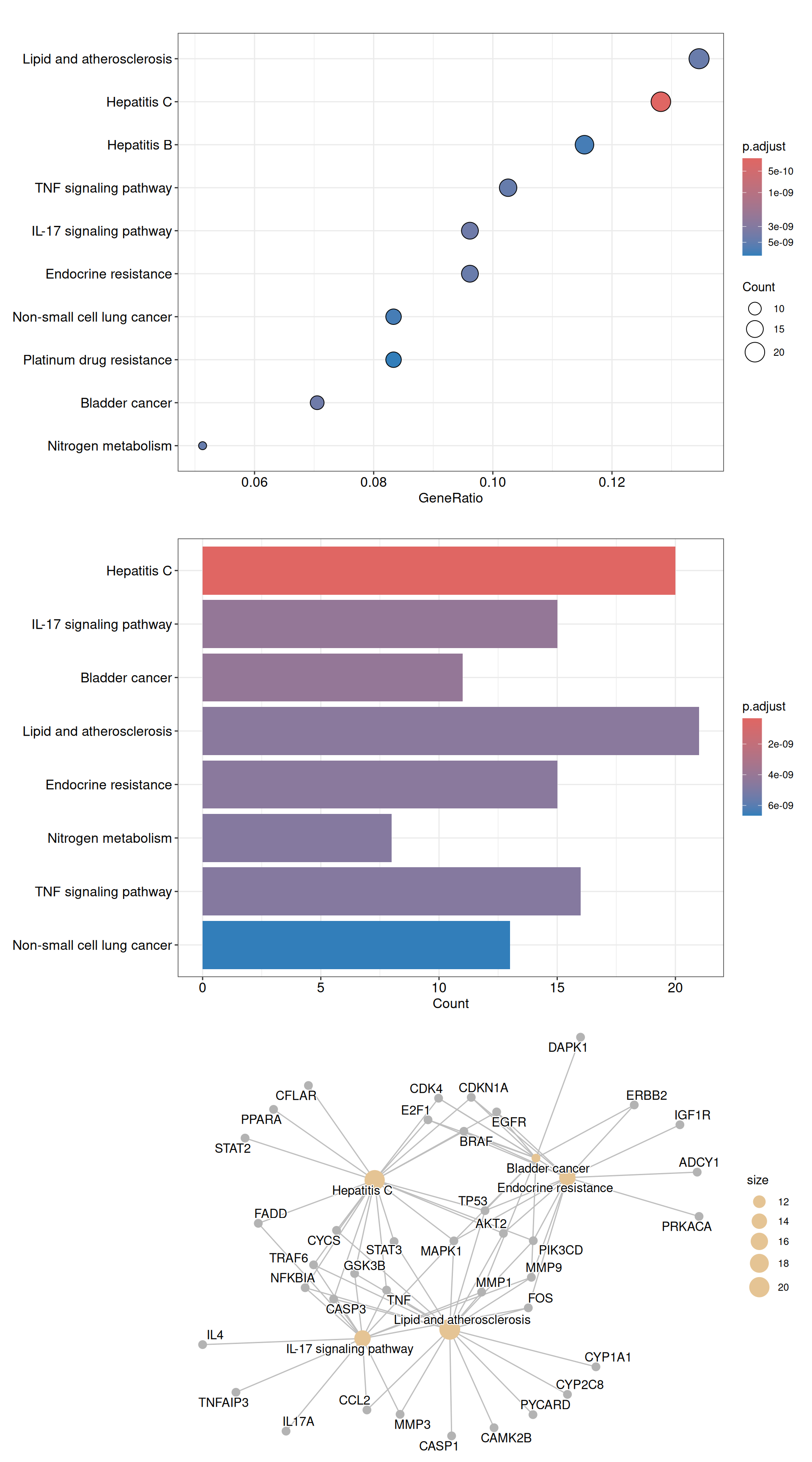
6.2 GO enrichment analysis
Gene Ontology (GO) is a widely used framework that provides a structured, controlled vocabulary for annotating gene functions. It consists of three orthogonal ontologies:
- BP (Biological Process): biological objectives to which a gene contributes (e.g., apoptosis, signal transduction).
- CC (Cellular Component): cellular locations where a gene product is active (e.g., nucleus, membrane).
- MF (Molecular Function): molecular activities of a gene product (e.g., kinase activity, DNA binding).
GO enrichment analysis tests whether GO terms are over-represented in a gene set relative to the genome background, thereby revealing the biological functions and cellular contexts associated with the input genes.
Here, we reused the gene set and Entrez IDs prepared in the KEGG section above. Unlike enrichKEGG(), enrichGO() also supports keyType = "SYMBOL" directly via the OrgDb annotation, but for consistency we continued to use Entrez IDs.
go_bp <- enrichGO(gene = gene_id$ENTREZID,
OrgDb = org.Hs.eg.db,
ont = "BP",
keyType = "ENTREZID",
readable = TRUE)
head(go_bp)#> ID Description GeneRatio
#> GO:0009410 GO:0009410 response to xenobiotic stimulus 31/176
#> GO:0043434 GO:0043434 response to peptide hormone 31/176
#> GO:0032868 GO:0032868 response to insulin 25/176
#> GO:0071375 GO:0071375 cellular response to peptide hormone stimulus 26/176
#> GO:0032496 GO:0032496 response to lipopolysaccharide 25/176
#> GO:0050727 GO:0050727 regulation of inflammatory response 27/176
#> BgRatio RichFactor FoldEnrichment zScore pvalue
#> GO:0009410 453/18805 0.06843267 7.311798 13.21743 3.003553e-18
#> GO:0043434 494/18805 0.06275304 6.704948 12.48952 3.519275e-17
#> GO:0032868 324/18805 0.07716049 8.244336 12.78483 4.542069e-16
#> GO:0071375 369/18805 0.07046070 7.528486 12.31059 1.007449e-15
#> GO:0032496 357/18805 0.07002801 7.482254 12.01911 4.333062e-15
#> GO:0050727 442/18805 0.06108597 6.526828 11.42883 9.112708e-15
#> p.adjust qvalue
#> GO:0009410 1.149760e-14 6.547745e-15
#> GO:0043434 6.735893e-14 3.836010e-14
#> GO:0032868 5.795680e-13 3.300570e-13
#> GO:0071375 9.641287e-13 5.490597e-13
#> GO:0032496 3.317392e-12 1.889215e-12
#> GO:0050727 5.813908e-12 3.310951e-12
#> geneID
#> GO:0009410 BCHE/CYP3A4/ADCY1/AOX1/ABL1/EGFR/ADAM17/TNF/GSTA1/NQO1/CDKN1A/TP53/EHMT2/BRAF/IL10/CASP3/PRKCB/CDK4/XPC/FBP1/CYP1A1/FOS/CAT/ADORA2A/CYP2C8/CA9/NOS2/TOP1/SREBF1/E2F1/CBR3
#> GO:0043434 STAT3/PRKACA/MAPK1/AGTR2/ERBB2/CA2/PKLR/EGFR/GSK3B/PKM/PPARA/INSR/PRKCD/GPT/ERBB4/BRAF/CTSD/IL10/LEP/NR1H4/PRKCB/MET/AKT2/FBP1/FOS/CFLAR/IGF1R/CAT/STAT2/NCOA2/SREBF1
#> GO:0032868 MAPK1/ERBB2/PKLR/EGFR/GSK3B/PKM/PPARA/INSR/PRKCD/GPT/ERBB4/CTSD/IL10/LEP/NR1H4/PRKCB/MET/AKT2/FBP1/FOS/CFLAR/IGF1R/CAT/NCOA2/SREBF1
#> GO:0071375 STAT3/PRKACA/MAPK1/AGTR2/ERBB2/CA2/PKLR/EGFR/GSK3B/PKM/INSR/PRKCD/GPT/ERBB4/CTSD/LEP/NR1H4/PRKCB/MET/AKT2/FBP1/FOS/CFLAR/IGF1R/NCOA2/SREBF1
#> GO:0032496 MMP3/MAPK1/ABL1/ADAM17/TNF/CTSG/NQO1/MAPKAPK2/TRAF6/XIAP/CASP1/MMP9/CCL2/IL10/NR1H4/CASP3/CDK4/CYP1A1/FOS/TNFAIP3/NOS2/PYCARD/PPARD/NFKBIA/E2F1
#> GO:0050727 YES1/STAT3/MMP3/SYK/IL4/PPARA/TNF/PRKCD/XIAP/PIK3CG/CASP1/MMP9/IL10/NR1H4/CASP3/FYN/PBK/ADORA1/PLA2G2D/TNFAIP3/ADORA2A/PYCARD/ALOX15/ADORA2B/IL17A/PPARD/NFKBIA
#> Count
#> GO:0009410 31
#> GO:0043434 31
#> GO:0032868 25
#> GO:0071375 26
#> GO:0032496 25
#> GO:0050727 27To perform enrichment for Cellular Component (CC), Molecular Function (MF), or all three categories simultaneously, simply change the ont parameter:
## Cellular Component (CC)
go_cc <- enrichGO(gene = gene_id$ENTREZID, OrgDb = org.Hs.eg.db,
ont = "CC", keyType = "ENTREZID", readable = TRUE)
## Molecular Function (MF)
go_mf <- enrichGO(gene = gene_id$ENTREZID, OrgDb = org.Hs.eg.db,
ont = "MF", keyType = "ENTREZID", readable = TRUE)
## All GO categories (ALL)
go_all <- enrichGO(gene = gene_id$ENTREZID, OrgDb = org.Hs.eg.db,
ont = "ALL", keyType = "ENTREZID", readable = TRUE)Similarly, GO enrichment results can be visualized with enrichplot (for further details, refer to the enrichplot documentation). For the combined ont = "ALL" result, a faceted bar plot provides a convenient comparison across BP, CC, and MF:
p_all <- barplot(
go_all,
x = "Count",
showCategory = 10,
split = "ONTOLOGY"
) +
aes(fill = ONTOLOGY) +
scale_fill_brewer(palette = "Set2") +
enrichplot::autofacet(by = "row", scales = "free")
p_all + scale_y_discrete()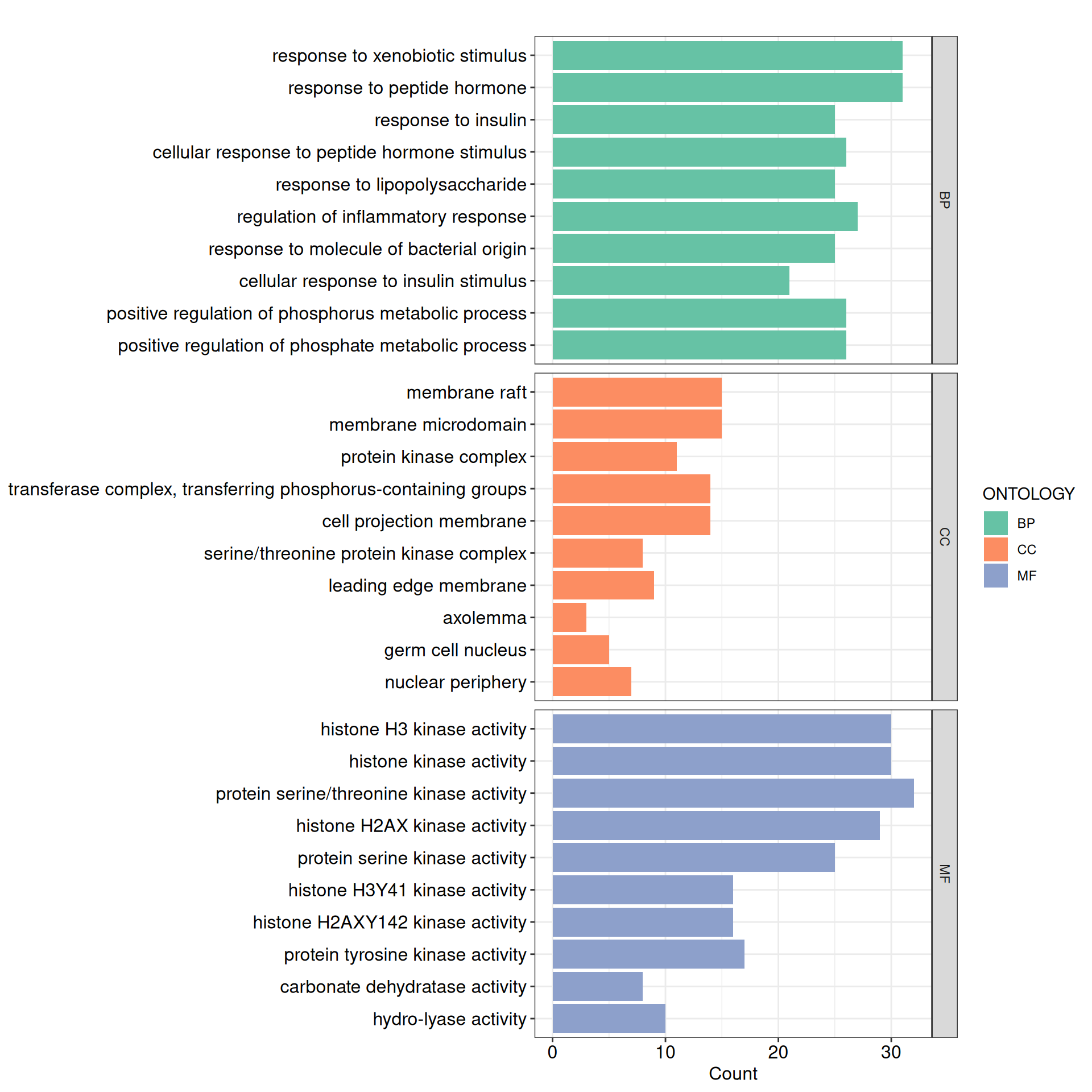
6.3 Advanced visualization
TCMDATA provides several publication-ready visualization functions tailored for enrichment results in network pharmacology research.
6.3.1 Lollipop plot
The lollipop plot offers an alternative to the standard dot/bar plot, combining line segments with terminal points to display enrichment significance and gene counts. gglollipop() is built on top of enrichplot and shares compatible grammar:
p_lollipop <- gglollipop(go_bp, line.col = "skyblue",
line.type = "dashed",
palette = "Spectral", top_n = 10) +
theme(strip.text.y = element_text(angle = 0))
p_lollipop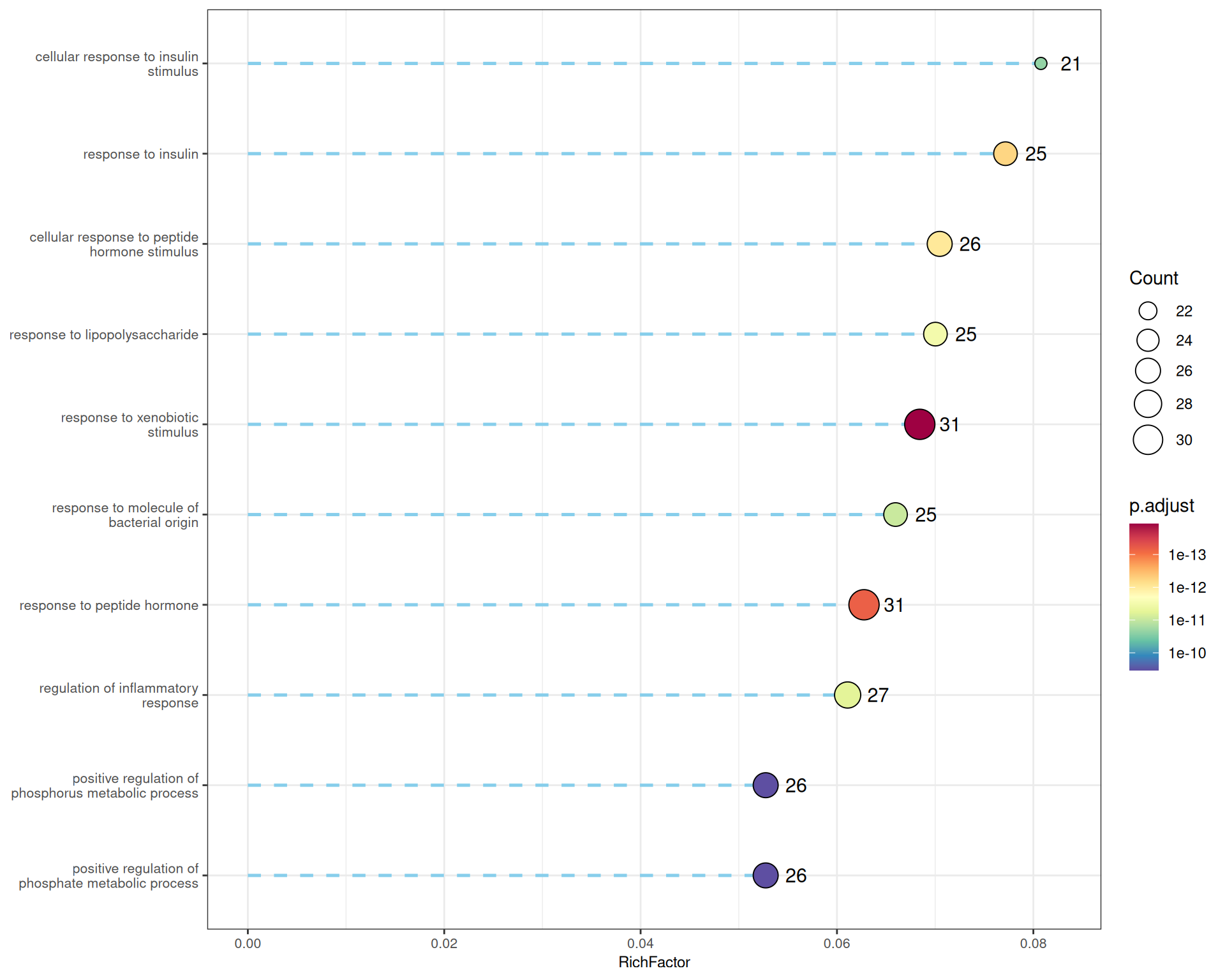
For the combined ont = "ALL" result, lollipop plots can be faceted by ontology to compare enriched terms across BP, CC, and MF:
p_lollipop_all <- gglollipop(go_all, line.col = "orange",
split = "ONTOLOGY",line.type = "dashed",
palette = "PiYG", top_n = 5, show_count = FALSE) +
facet_grid(ONTOLOGY ~ ., scales = "free_y") +
theme(strip.text.y = element_text(angle = 0))
p_lollipop_all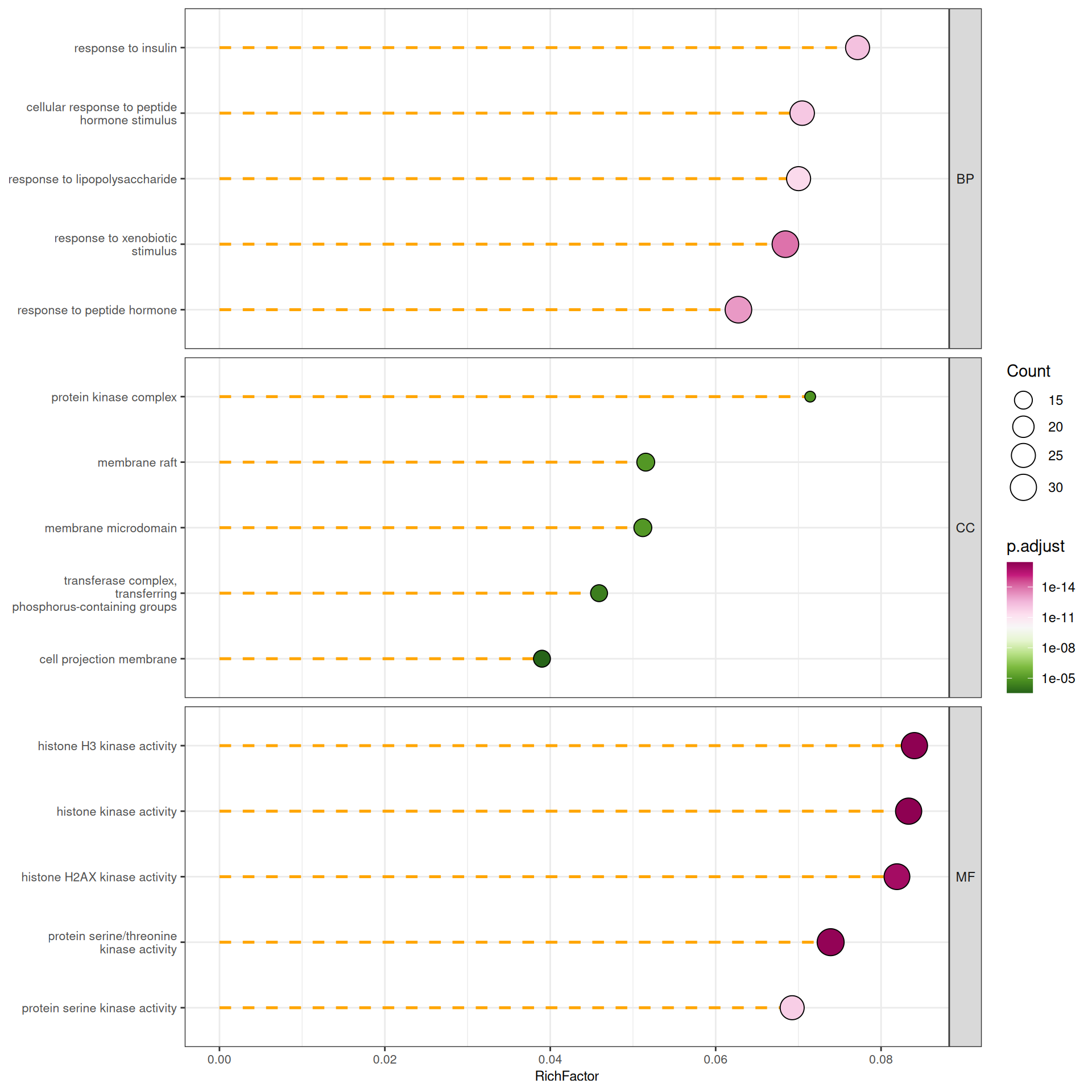
6.3.2 Dot-sankey plot
The dot-sankey plot combines a Sankey diagram (showing gene–pathway alluvial flows) with a dot plot (encoding significance and gene count) in a single panel. This dual-view layout facilitates simultaneous inspection of which genes contribute to each pathway and how significant each enrichment is:
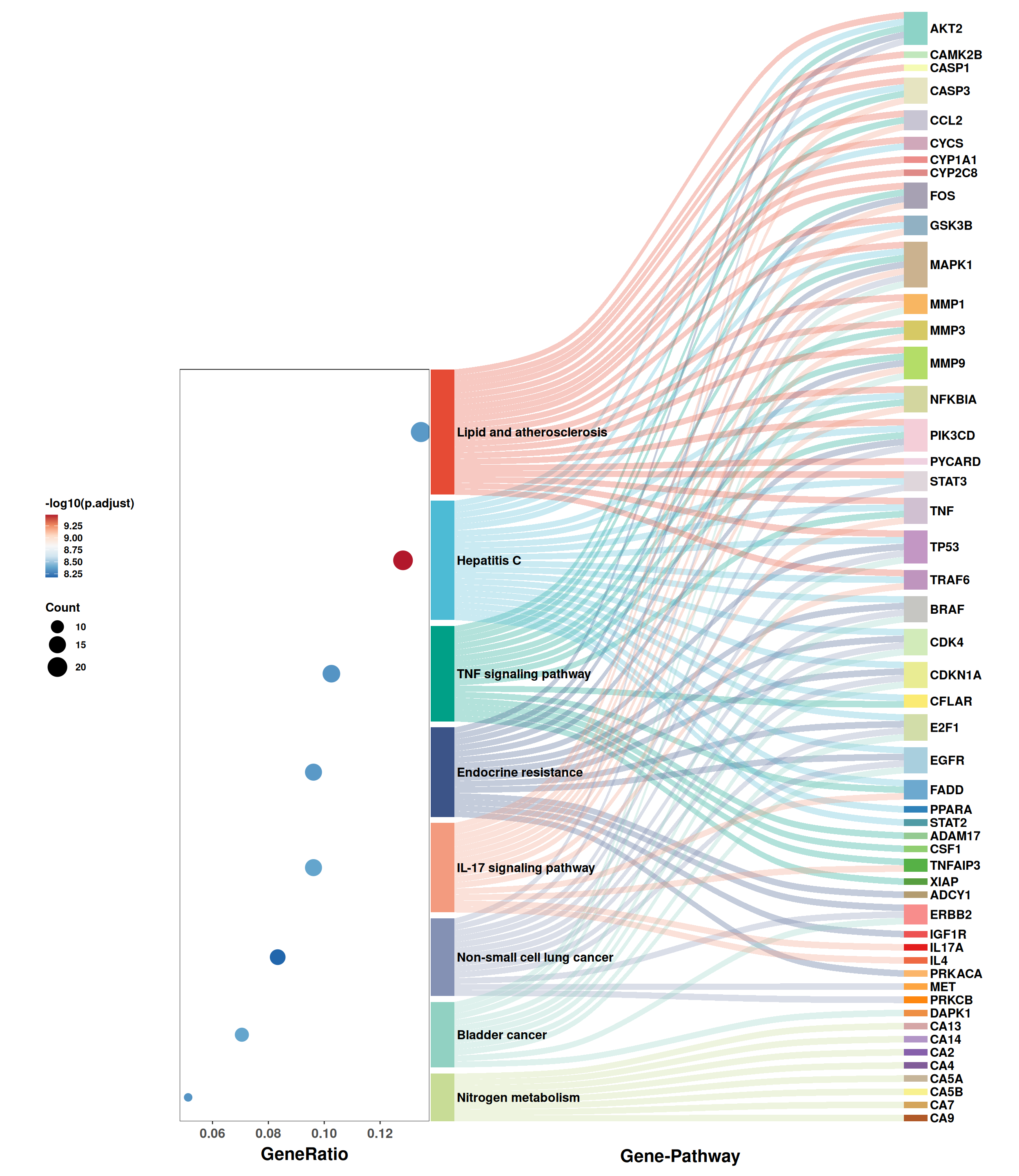
6.3.3 GO bar plot
go_barplot() provides a colored bar chart specifically designed for enrichGO(ont = "ALL") results, where bars are grouped and colored by ontology category (BP, CC, MF). These figures are commonly used in network pharmacology publications for presenting GO enrichment results:
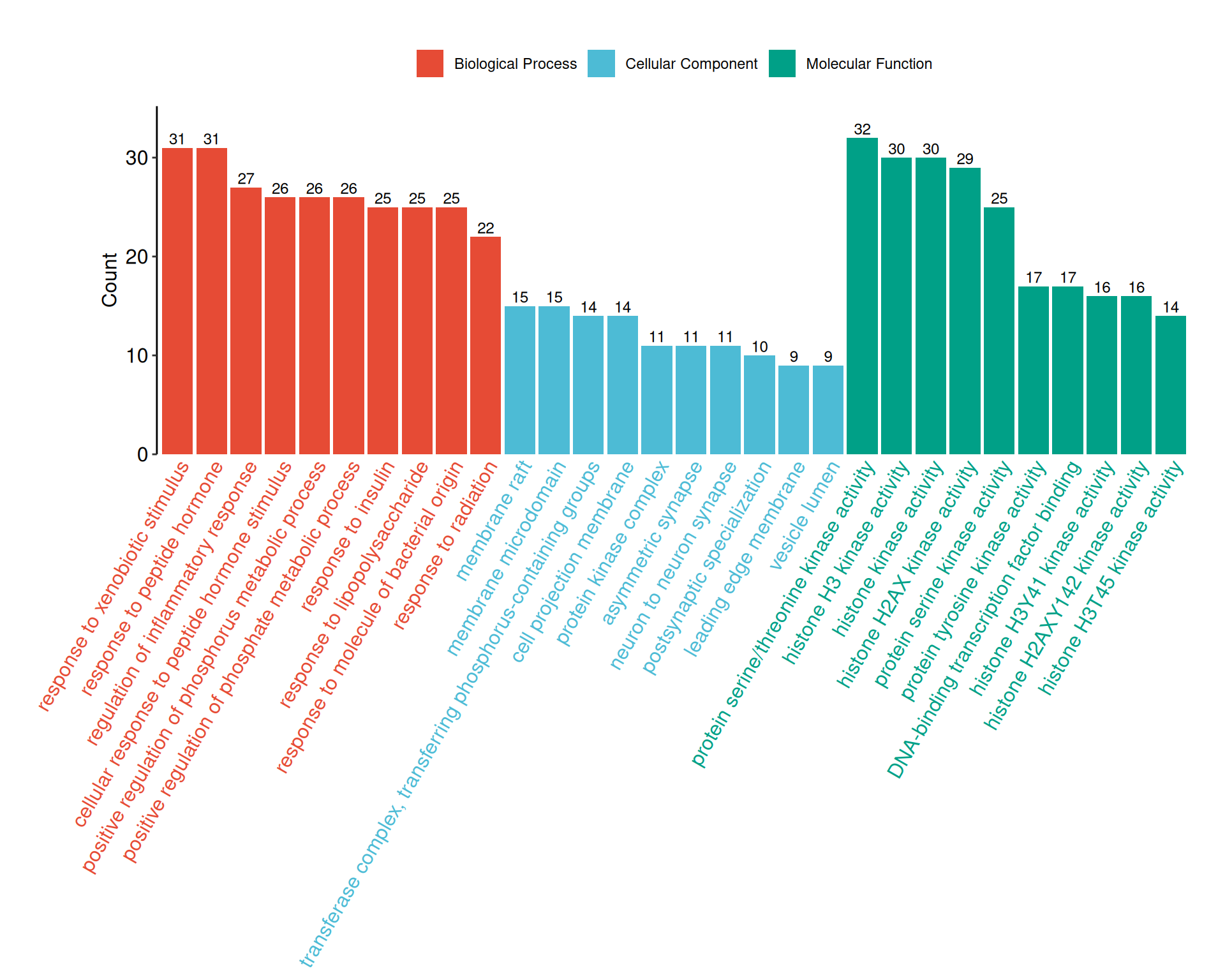
6.4 Gene Set Enrichment Analysis (GSEA)
Unlike ORA, which requires a pre-defined list of significant genes, Gene Set Enrichment Analysis (GSEA) uses the entire ranked gene list to detect coordinated changes in gene sets. This approach avoids arbitrary significance cutoffs and is more sensitive to subtle but consistent expression shifts across a pathway.
The typical GSEA workflow requires a named numeric vector (e.g., log2 fold change or Wald statistic), sorted in decreasing order, with gene identifiers as names. Here, we use the built-in deg_earlydn dataset from TCMDATA to construct such a ranked list.
6.4.1 Prepare ranked gene list
data(deg_earlydn)
## convert gene symbols to Entrez IDs
deg_ids <- bitr(deg_earlydn$names,
fromType = "SYMBOL",
toType = "ENTREZID",
OrgDb = org.Hs.eg.db)
## merge with DESeq2 results and build ranked list
deg_merged <- merge(deg_earlydn, deg_ids, by.x = "names", by.y = "SYMBOL")
gene_rank <- sort(setNames(deg_merged$log2FoldChange, deg_merged$ENTREZID),
decreasing = TRUE)
head(gene_rank)#> 9370 100874095 1149 4633 4625 100189286
#> 8.517210 7.070620 5.368624 5.256895 4.874813 4.7938946.4.2 KEGG GSEA analysis
gsea_kegg <- gseKEGG(geneList = gene_rank,
organism = "hsa",
eps = 0,
pvalueCutoff = 0.05)
gsea_kegg <- setReadable(gsea_kegg, OrgDb = org.Hs.eg.db, keyType = "ENTREZID")
head(gsea_kegg)#> ID Description setSize
#> hsa00970 hsa00970 Aminoacyl-tRNA biosynthesis 136
#> hsa04380 hsa04380 Osteoclast differentiation 132
#> hsa04668 hsa04668 TNF signaling pathway 117
#> hsa04657 hsa04657 IL-17 signaling pathway 82
#> hsa05323 hsa05323 Rheumatoid arthritis 88
#> hsa04060 hsa04060 Cytokine-cytokine receptor interaction 244
#> enrichmentScore NES pvalue p.adjust qvalue rank
#> hsa00970 0.7812747 3.088860 2.452704e-30 8.633517e-28 6.093032e-28 734
#> hsa04380 -0.6513337 -2.546759 1.345775e-14 2.368564e-12 1.671594e-12 2855
#> hsa04668 -0.6777418 -2.604701 4.909992e-14 5.761058e-12 4.065818e-12 1129
#> hsa04657 -0.7364961 -2.699637 7.287049e-14 6.234807e-12 4.400163e-12 1767
#> hsa05323 -0.7173842 -2.648602 8.856260e-14 6.234807e-12 4.400163e-12 2438
#> hsa04060 -0.5366194 -2.299662 7.115359e-13 4.174344e-11 2.946008e-11 2557
#> leading_edge
#> hsa00970 tags=32%, list=3%, signal=32%
#> hsa04380 tags=25%, list=11%, signal=22%
#> hsa04668 tags=21%, list=4%, signal=20%
#> hsa04657 tags=30%, list=6%, signal=29%
#> hsa05323 tags=34%, list=9%, signal=31%
#> hsa04060 tags=25%, list=9%, signal=23%
#> core_enrichment
#> hsa00970 TRP-TGG1-1/TRR-CCG2-1/TRH-GTG1-8/TRK-TTT3-1/TRH-GTG1-9/TRE-TTC2-1/TRR-CCT3-1/TRS-GCT6-1/TRR-TCG1-1/TRP-AGG2-6/TRR-TCG3-1/TRF-GAA2-1/TRH-GTG1-7/TRL-TAG2-1/TRQ-CTG6-1/TRK-TTT5-1/TRY-GTA2-1/TRL-TAG3-1/TRY-GTA5-1/TRL-CAG2-1/TRC-GCA11-1/TRL-CAA1-1/TRI-AAT5-5/TRE-CTC1-3/TRE-CTC1-4/TRI-AAT2-1/TRC-GCA8-1/TRE-CTC1-5/TRG-GCC2-4/TRC-GCA2-4/TRG-GCC1-5/TRY-GTA5-5/TRD-GTC2-10/TRT-AGT1-1/TRR-ACG1-2/TRA-TGC4-1/TRG-TCC2-6/TRS-AGA2-2/TRV-CAC1-5/TRK-CTT2-2/TRK-CTT3-1/TRS-GCT3-1/TRC-GCA3-1/TRG-GCC3-1
#> hsa04380 PLCG2/CALCR/MAP3K14/IFNGR1/PPP3R1/FCGR2A/LILRB2/PIK3CA/FCGR3A/TGFB2/FOSL1/NFKBIA/NFATC1/IL1A/LILRA6/LILRB4/RELB/TNFSF11/LILRB3/TREM2/NFKB2/OSCAR/JUND/PPP3R2/FOSL2/FCGR3B/IFNG/IL1B/JUN/SOCS3/JUNB/FOS/FOSB
#> hsa04668 CXCL6/VCAM1/IL18R1/CEBPB/TRAF1/ICAM1/TNFAIP3/CCL2/BCL3/MMP9/BIRC3/LIF/PTGS2/CXCL1/CXCL3/IL1B/SELE/JUN/SOCS3/CCL20/JUNB/CXCL2/IL6/FOS
#> hsa04657 MUC5AC/HSP90AA1/FOSL1/NFKBIA/CXCL6/CEBPB/CSF3/TNFAIP3/S100A9/JUND/CCL2/MMP9/PTGS2/CXCL1/S100A8/CXCL3/IFNG/CXCL8/IL1B/JUN/CCL20/CXCL2/IL6/FOS/FOSB
#> hsa05323 ITGB2/IL11/VEGFA/TGFB3/ATP6V1G3/ATP6V0D2/ATP6V1D/TLR2/TGFB2/IL1A/FLT1/IL18/CXCL6/TNFSF11/CD86/ICAM1/CCL2/CXCL1/CTLA4/CXCL3/IFNG/CCL3L3/CXCL8/IL1B/JUN/CCL20/CXCL2/IL6/CCL3/FOS
#> hsa04060 OSMR/IFNGR1/GDF3/IL10/CX3CL1/TNFRSF10B/IL11/TGFB3/CCL16/IL2RG/RELT/CRLF2/TGFB2/TNFRSF12A/IL24/TNFRSF9/IL1A/CLCF1/IL18/CCL18/CD70/CCL23/CCR7/CXCL6/IL1RL1/TNFSF11/TNFSF10/PPBP/IL18R1/IL18RAP/CSF3R/CCR3/CCL22/IL7R/TNFRSF13C/CSF3/CCL2/CXCR1/CXCR4/CXCR2/PF4V1/LIF/TNFRSF13B/CXCL1/IL1RN/CCL8/CXCL13/CXCL3/IFNG/CCL3L3/INHBB/TNFSF9/CCL4/OSM/CXCL8/IL1B/GDF15/CCL20/CCL4L2/CXCL2/IL6/CCL3The classic GSEA running-score plot visualizes the enrichment profile of selected pathways. gseaplot2() from enrichplot supports displaying multiple gene sets simultaneously:
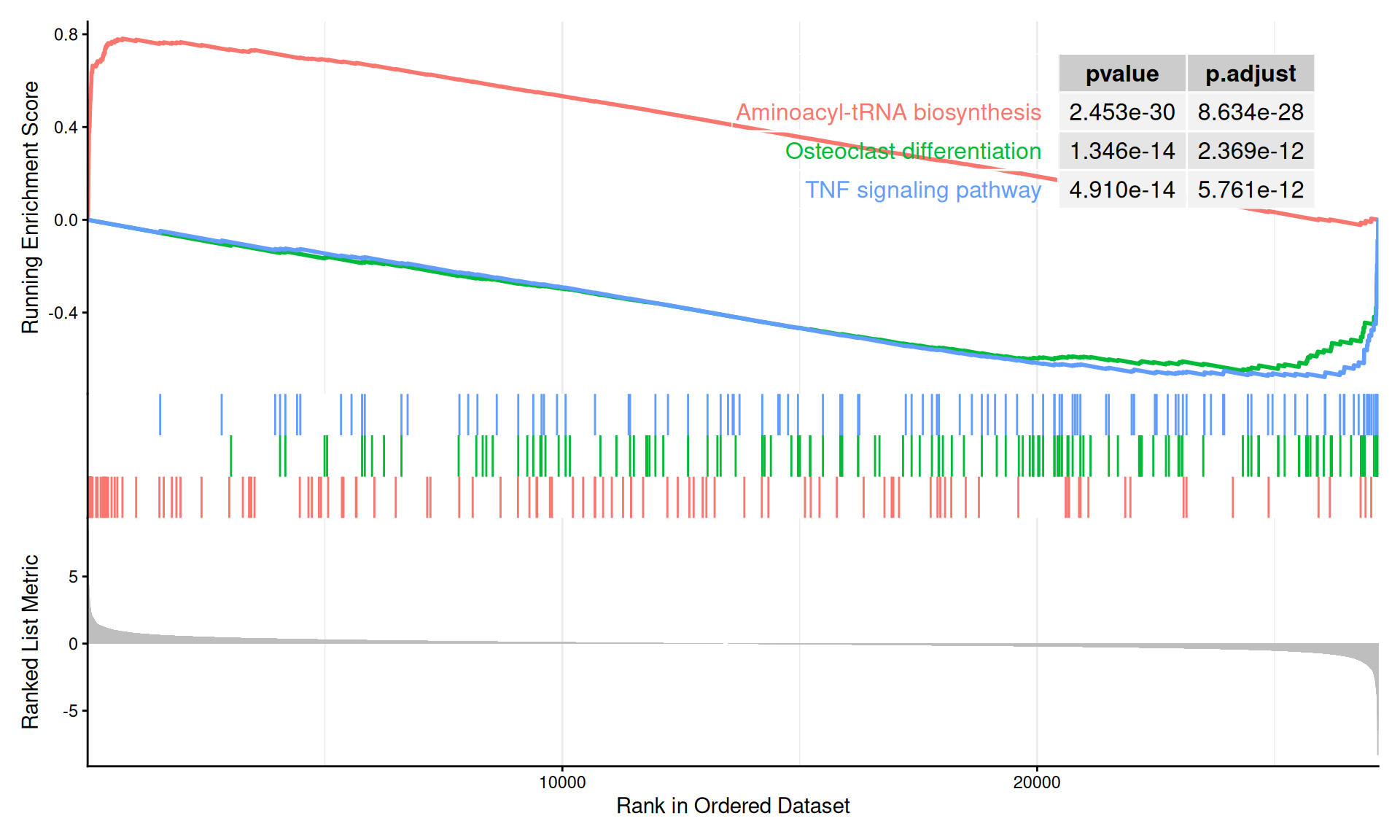
6.4.3 GO GSEA analysis
gsea_go <- gseGO(geneList = gene_rank,
OrgDb = org.Hs.eg.db,
ont = "BP",
eps = 0,
pvalueCutoff = 0.05)
gsea_go <- setReadable(gsea_go, OrgDb = org.Hs.eg.db, keyType = "ENTREZID")
head(gsea_go)#> ID
#> GO:0002460 GO:0002460
#> GO:0002443 GO:0002443
#> GO:0002449 GO:0002449
#> GO:0019724 GO:0019724
#> GO:0016064 GO:0016064
#> GO:0035195 GO:0035195
#> Description
#> GO:0002460 adaptive immune response based on somatic recombination of immune receptors built from immunoglobulin superfamily domains
#> GO:0002443 leukocyte mediated immunity
#> GO:0002449 lymphocyte mediated immunity
#> GO:0019724 B cell mediated immunity
#> GO:0016064 immunoglobulin mediated immune response
#> GO:0035195 miRNA-mediated post-transcriptional gene silencing
#> setSize enrichmentScore NES pvalue p.adjust
#> GO:0002460 367 -0.5962006 -2.649126 7.191799e-27 4.421518e-23
#> GO:0002443 446 -0.5540426 -2.507669 2.307337e-25 7.092755e-22
#> GO:0002449 352 -0.5705852 -2.526617 2.865721e-21 5.872818e-18
#> GO:0019724 187 -0.6650901 -2.711805 1.080066e-20 1.660061e-17
#> GO:0016064 184 -0.6704404 -2.728365 1.781633e-20 2.190697e-17
#> GO:0035195 296 0.5648152 2.477770 4.576711e-19 4.689603e-16
#> qvalue rank leading_edge
#> GO:0002460 3.618611e-23 3405 tags=34%, list=13%, signal=30%
#> GO:0002443 5.804775e-22 3899 tags=34%, list=14%, signal=30%
#> GO:0002449 4.806367e-18 3144 tags=35%, list=12%, signal=32%
#> GO:0019724 1.358609e-17 2665 tags=40%, list=10%, signal=36%
#> GO:0016064 1.792886e-17 2665 tags=40%, list=10%, signal=37%
#> GO:0035195 3.838014e-16 1814 tags=31%, list=7%, signal=30%
#> core_enrichment
#> GO:0002460 IL23R/IL6R/ZBTB1/STAT4/EXO1/SLC11A1/C5/SERPING1/GATA3/ANXA1/CR1/CD1E/WAS/MYO1G/MEF2C/TLR8/SLAMF6/TNFRSF1B/FCER1G/KLRC1/CD19/LIG4/PRF1/CEACAM1/HSPD1/IL10/IGHV3-66/CD7/SLAMF1/SCART1/FCMR/FCGR2A/IGHV4-61/CLU/BCL6/TFRC/FCGR3A/IGLL5/IGLC1/HLX/PTPRC/C9/IGHV3-73/ARID5A/IGHV3-43/KMT5C/IGHV1-18/CLCF1/IL18/MIR21/CD70/LILRB4/BACH2/IGHV3-48/IGHV3-15/IGHG4/RELB/IL1RL1/IGKC/C3/LY9/KLHL6/TREM2/CFI/IGHV5-10-1/IGHV3-49/IGHV4-39/IL18R1/NFKB2/CD1A/IGHV1-69/IGHV3-74/IGHV7-4-1/IGLC6/IGHV6-1/IGHE/CD55/IGLC2/IGHV1-3/CLC/IGHV3-11/ICAM1/IL7R/ULBP1/IGHV3-64D/IGHG2/KLRD1/IGHG1/IGHV1-69D/IGHG3/IGHV3-23/TNFAIP3/IGHV1-58/CD1B/IGHA1/C2/IGHV3-16/IGLC3/BCL3/FCGR3B/IGHD/IGHV3-13/ZC3H12A/IGHA2/IGHV3-7/NFKBID/PSG9/CR2/IGHV3-53/NFKBIZ/IGHV3-33/IGHV5-51/IGHV3-20/IGHV3-30/IGLC7/IGHV3-72/CXCL13/CD69/IGHV4-28/IL1B/SOCS3/JUNB/IGHV4-4/IL6
#> GO:0002443 CD46/CD84/PTAFR/IL2RB/NKG7/NOS2/IL13RA2/NBN/CLEC7A/RNF19B/CYRIB/NLRP3/SCNN1B/IL23R/IL6R/ZBTB1/EXO1/RAC2/DHX36/C5/SERPING1/GATA3/CR1/KLRB1/CD1E/WAS/MYO1G/PLCG2/TLR8/SLAMF6/GZMB/VAMP7/TNFRSF1B/KLRC2/FCER1G/KLRC1/STXBP1/FGR/CD19/LIG4/PRF1/RAB44/PIK3R6/CEACAM1/HSPD1/ITGB2/IL10/IGHV3-66/CD7/TICAM1/SLAMF1/SCART1/SERPINB9/ADGRE2/ARL8B/STXBP3/FCMR/FCGR2A/ANXA3/IGHV4-61/CD96/KIT/CLU/BCL6/TFRC/FCGR3A/IGLL5/IGLC1/PTPRC/C9/IGHV3-73/ARID5A/IGHV3-43/DNASE1L3/KMT5C/SLAMF7/IGHV1-18/CLCF1/SLC18A2/IL18/CLNK/CD70/LILRB4/IGHV3-48/IGHV3-15/IGHG4/CXCL6/IGKC/C3/TREM2/CFI/IGHV5-10-1/STAP1/IGHV3-49/NCR1/IGHV4-39/IL18R1/CD1A/IGHV1-69/IL18RAP/SH2D1B/IGHV3-74/IGHV7-4-1/IGLC6/IGHV6-1/IGHE/CD55/IGLC2/CD177/IGHV1-3/CLC/IGHV3-11/ICAM1/KLRC3/IL7R/ULBP1/IGHV3-64D/IGHG2/KLRD1/IGHG1/IGHV1-69D/IGHG3/IGHV3-23/IGHV1-58/CD1B/FOXF1/IGHA1/C2/IGHV3-16/IGLC3/BCL3/FCGR3B/IGHD/IGHV3-13/IGHA2/IGHV3-7/CR2/IGHV3-53/NFKBIZ/IGHV3-33/IGHV5-51/IGHV3-20/IGHV3-30/IGLC7/IGHV3-72/FCAR/IGHV4-28/TREM1/IL1B/IGHV4-4/IL6/CCL3/NR4A3
#> GO:0002449 IL2RB/NKG7/IL13RA2/NBN/RNF19B/CYRIB/NLRP3/IL23R/ZBTB1/EXO1/C5/SERPING1/GATA3/CR1/KLRB1/CD1E/WAS/MYO1G/TLR8/SLAMF6/GZMB/VAMP7/TNFRSF1B/KLRC2/FCER1G/KLRC1/CD19/LIG4/PRF1/PIK3R6/CEACAM1/HSPD1/IL10/IGHV3-66/CD7/SLAMF1/SCART1/SERPINB9/ARL8B/FCMR/FCGR2A/IGHV4-61/CD96/CLU/BCL6/TFRC/FCGR3A/IGLL5/IGLC1/PTPRC/C9/IGHV3-73/ARID5A/IGHV3-43/KMT5C/SLAMF7/IGHV1-18/CLCF1/IL18/CLNK/CD70/LILRB4/IGHV3-48/IGHV3-15/IGHG4/IGKC/C3/TREM2/CFI/IGHV5-10-1/IGHV3-49/NCR1/IGHV4-39/IL18R1/CD1A/IGHV1-69/IL18RAP/SH2D1B/IGHV3-74/IGHV7-4-1/IGLC6/IGHV6-1/IGHE/CD55/IGLC2/IGHV1-3/CLC/IGHV3-11/ICAM1/KLRC3/IL7R/ULBP1/IGHV3-64D/IGHG2/KLRD1/IGHG1/IGHV1-69D/IGHG3/IGHV3-23/IGHV1-58/CD1B/IGHA1/C2/IGHV3-16/IGLC3/BCL3/FCGR3B/IGHD/IGHV3-13/IGHA2/IGHV3-7/CR2/IGHV3-53/NFKBIZ/IGHV3-33/IGHV5-51/IGHV3-20/IGHV3-30/IGLC7/IGHV3-72/IGHV4-28/IL1B/IGHV4-4/IL6
#> GO:0019724 C5/SERPING1/CR1/TLR8/FCER1G/CD19/LIG4/HSPD1/IL10/IGHV3-66/FCMR/FCGR2A/IGHV4-61/CLU/BCL6/TFRC/FCGR3A/IGLL5/IGLC1/PTPRC/C9/IGHV3-73/IGHV3-43/KMT5C/IGHV1-18/CLCF1/CD70/IGHV3-48/IGHV3-15/IGHG4/IGKC/C3/TREM2/CFI/IGHV5-10-1/IGHV3-49/IGHV4-39/IGHV1-69/IGHV3-74/IGHV7-4-1/IGLC6/IGHV6-1/IGHE/CD55/IGLC2/IGHV1-3/IGHV3-11/IGHV3-64D/IGHG2/IGHG1/IGHV1-69D/IGHG3/IGHV3-23/IGHV1-58/IGHA1/C2/IGHV3-16/IGLC3/BCL3/FCGR3B/IGHD/IGHV3-13/IGHA2/IGHV3-7/CR2/IGHV3-53/NFKBIZ/IGHV3-33/IGHV5-51/IGHV3-20/IGHV3-30/IGLC7/IGHV3-72/IGHV4-28/IGHV4-4
#> GO:0016064 C5/SERPING1/CR1/TLR8/FCER1G/CD19/LIG4/HSPD1/IL10/IGHV3-66/FCMR/FCGR2A/IGHV4-61/CLU/BCL6/TFRC/FCGR3A/IGLL5/IGLC1/PTPRC/C9/IGHV3-73/IGHV3-43/KMT5C/IGHV1-18/CLCF1/IGHV3-48/IGHV3-15/IGHG4/IGKC/C3/TREM2/CFI/IGHV5-10-1/IGHV3-49/IGHV4-39/IGHV1-69/IGHV3-74/IGHV7-4-1/IGLC6/IGHV6-1/IGHE/CD55/IGLC2/IGHV1-3/IGHV3-11/IGHV3-64D/IGHG2/IGHG1/IGHV1-69D/IGHG3/IGHV3-23/IGHV1-58/IGHA1/C2/IGHV3-16/IGLC3/BCL3/FCGR3B/IGHD/IGHV3-13/IGHA2/IGHV3-7/CR2/IGHV3-53/NFKBIZ/IGHV3-33/IGHV5-51/IGHV3-20/IGHV3-30/IGLC7/IGHV3-72/IGHV4-28/IGHV4-4
#> GO:0035195 MIR30D/MIR29A/MIRLET7G/MIR181B1/MIR362/MIR503/MIR425/MIR195/MIR103A2/MIRLET7A3/MIRLET7A1/MIRLET7B/MIRLET7D/MIR708/MIR148B/MIR99A/MIR345/MIRLET7C/MIR130A/MIR27B/MIR30B/MIR193A/MIR30C1/MIR107/MIR190A/MIR29B2/MIR501/MIR331/MIR23B/MIR326/MIR1251/MIR125B2/MIR98/MIR204/MIR532/MIR126/MIR3184/MIR24-1/MIR497/MIR188/MIRLET7F2/MIR29C/LIN28A/MIR455/MIRLET7F1/MIR101-1/MIRLET7I/MIR106B/MIR339/MIR191/MIR194-2/MIR93/MIR3074/MIR135A2/MIR30E/MIR500A/MIR196B/MIR152/MIR187/MIR125B1/MIR575/MIR10A/MIR224/MIR143/MIR219A1/MIR135B/MIRLET7A2/MIR500B/MIR25/MIR657/MIR574/MIR200B/MIR185/MIR378A/MIR3619/MIR214/MIRLET7E/MIR423/MIR26A1/MIR197/MIR28/MIR9-2HG/MIR615/MIR3120/MIR940/MIR26B/MIR1306/MIR1207/MIR192/MIR1225/MIR10B/MIR589/MIR365AA ridge plot provides a density overview of enrichment score distributions across top GO terms:
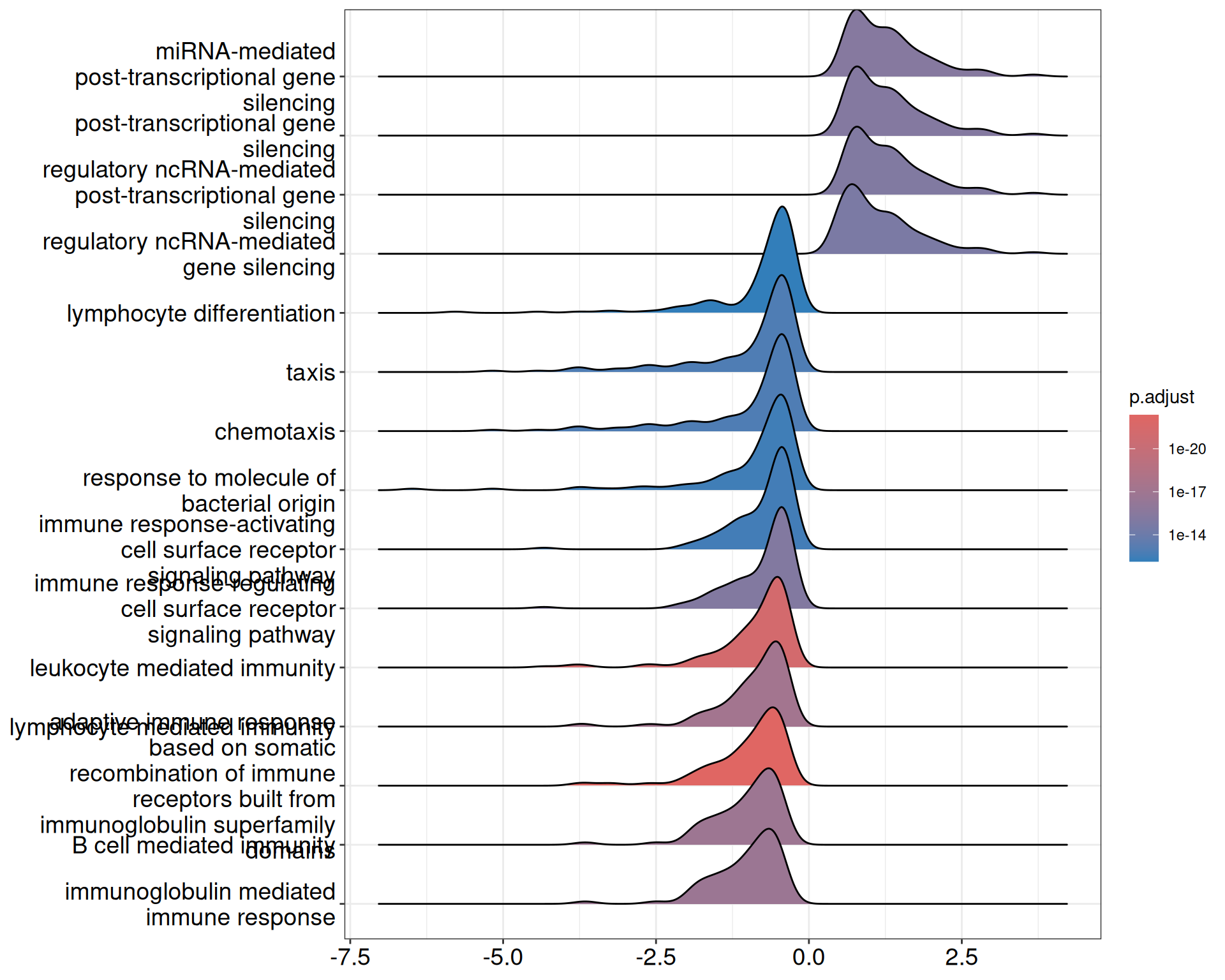
6.5 Session information
#> R version 4.5.3 (2026-03-11)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
#> [3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
#> [5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
#> [7] LC_PAPER=en_US.UTF-8 LC_NAME=C
#> [9] LC_ADDRESS=C LC_TELEPHONE=C
#> [11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats4 stats graphics grDevices utils datasets methods
#> [8] base
#>
#> other attached packages:
#> [1] org.Hs.eg.db_3.22.0 AnnotationDbi_1.72.0 IRanges_2.44.0
#> [4] S4Vectors_0.48.1 Biobase_2.70.0 BiocGenerics_0.56.0
#> [7] generics_0.1.4 clusterProfiler_4.18.4 aplot_0.2.9
#> [10] ggrepel_0.9.8 ggtangle_0.1.1 igraph_2.2.3
#> [13] ggplot2_4.0.2 dplyr_1.2.1 ivolcano_0.0.5
#> [16] enrichplot_1.30.5 TCMDATA_0.1.0
#>
#> loaded via a namespace (and not attached):
#> [1] RColorBrewer_1.1-3 rstudioapi_0.18.0 jsonlite_2.0.0
#> [4] shape_1.4.6.1 tidydr_0.0.6 magrittr_2.0.5
#> [7] farver_2.1.2 rmarkdown_2.31 GlobalOptions_0.1.4
#> [10] fs_2.0.1 vctrs_0.7.3 memoise_2.0.1
#> [13] ggtree_4.0.5 htmltools_0.5.9 gridGraphics_0.5-1
#> [16] sass_0.4.10 bslib_0.10.0 htmlwidgets_1.6.4
#> [19] plyr_1.8.9 cachem_1.1.0 lifecycle_1.0.5
#> [22] iterators_1.0.14 pkgconfig_2.0.3 Matrix_1.7-5
#> [25] R6_2.6.1 fastmap_1.2.0 gson_0.1.0
#> [28] clue_0.3-68 digest_0.6.39 colorspace_2.1-2
#> [31] ggnewscale_0.5.2 patchwork_1.3.2 RSQLite_2.4.6
#> [34] labeling_0.4.3 httr_1.4.8 polyclip_1.10-7
#> [37] compiler_4.5.3 bit64_4.6.0-1 fontquiver_0.2.1
#> [40] withr_3.0.2 doParallel_1.0.17 S7_0.2.1
#> [43] BiocParallel_1.44.0 DBI_1.3.0 ggforce_0.5.0
#> [46] R.utils_2.13.0 MASS_7.3-65 rappdirs_0.3.4
#> [49] rjson_0.2.23 tools_4.5.3 ape_5.8-1
#> [52] scatterpie_0.2.6 R.oo_1.27.1 glue_1.8.0
#> [55] nlme_3.1-168 GOSemSim_2.36.0 grid_4.5.3
#> [58] ggvenn_0.1.19 cluster_2.1.8.2 reshape2_1.4.5
#> [61] fgsea_1.36.2 gtable_0.3.6 R.methodsS3_1.8.2
#> [64] tidyr_1.3.2 data.table_1.18.2.1 XVector_0.50.0
#> [67] foreach_1.5.2 pillar_1.11.1 stringr_1.6.0
#> [70] yulab.utils_0.2.4 circlize_0.4.18 splines_4.5.3
#> [73] tweenr_2.0.3 treeio_1.34.0 lattice_0.22-9
#> [76] bit_4.6.0 tidyselect_1.2.1 fontLiberation_0.1.0
#> [79] GO.db_3.22.0 ComplexHeatmap_2.26.1 Biostrings_2.78.0
#> [82] knitr_1.51 gridExtra_2.3 fontBitstreamVera_0.1.1
#> [85] bookdown_0.46 Seqinfo_1.0.0 xfun_0.57
#> [88] matrixStats_1.5.0 stringi_1.8.7 lazyeval_0.2.3
#> [91] ggfun_0.2.0 yaml_2.3.12 evaluate_1.0.5
#> [94] codetools_0.2-20 gdtools_0.5.0 tibble_3.3.1
#> [97] qvalue_2.42.0 ggplotify_0.1.3 cli_3.6.6
#> [100] systemfonts_1.3.2 jquerylib_0.1.4 Rcpp_1.1.1
#> [103] png_0.1-9 parallel_4.5.3 blob_1.3.0
#> [106] ggalluvial_0.12.6 DOSE_4.4.0 tidytree_0.4.7
#> [109] ggiraph_0.9.6 ggridges_0.5.7 scales_1.4.0
#> [112] purrr_1.2.2 crayon_1.5.3 GetoptLong_1.1.1
#> [115] rlang_1.2.0 cowplot_1.2.0 fastmatch_1.1-8
#> [118] KEGGREST_1.50.0